INTRODUCTION
Plague is an extremely dangerous zoonotic infectious disease with natural foci. The etiological agent of plague is the gram-negative bacterium Yersinia pestis. Plague has left an unparalleled mark on human history, and it still retains its pandemic potential thousands of years later and can lead to public health emergencies. The species Y. pestis involves several main phylogenetic lineages, including the medieval biovar 2.MED (Medievalia) – the causative agent in the plague foci in the Russian Federation (RF) and neighboring countries, covering a total area of 1,959,965 km2 [1, 2]. The strains of this biovar exhibit ecological plasticity. Their populations exist in natural foci of various types, such as desert, semi-desert, mountain, high-mountain, and plain areas, including those with a high degree of aridity where other biovars and subspecies of Y. pestis do not circulate [1-5]. According to historical data, in the 20th century, Y. pestis was most active in the Caspian region foci, causing highly lethal plague outbreaks in the first half of this century [5]. Using the genome-wide SNP genotyping and phylogenetic reconstruction of 38 strains of the medieval Y. pestis biovar, we found that these plague outbreaks in the northern part of the Caspian region were caused by the strains of Y. pestis from the phylogroup 2.MED1 and a previously unknown phylogroup 2.MED4 [3, 4]. Comparison of the core genomes of these phylogroups revealed differences in their SNP profiles: 2.MED1 contained 14 SNPs, whereas 2.MED4 contained 9 SNPs. The unique polymorphisms in the core genome that differentiate the new phylogroup were found in the genes encoding biochemical processes. Our study aimed to conduct a comparative analysis of the phenotypic and genotypic characteristics of Y. pestis strains from the phylogroups 2.MED1 and 2.MED4, which caused large plague outbreaks in the Northern Caspian region in the first half of the 20th century.
MATERIALS AND METHODS
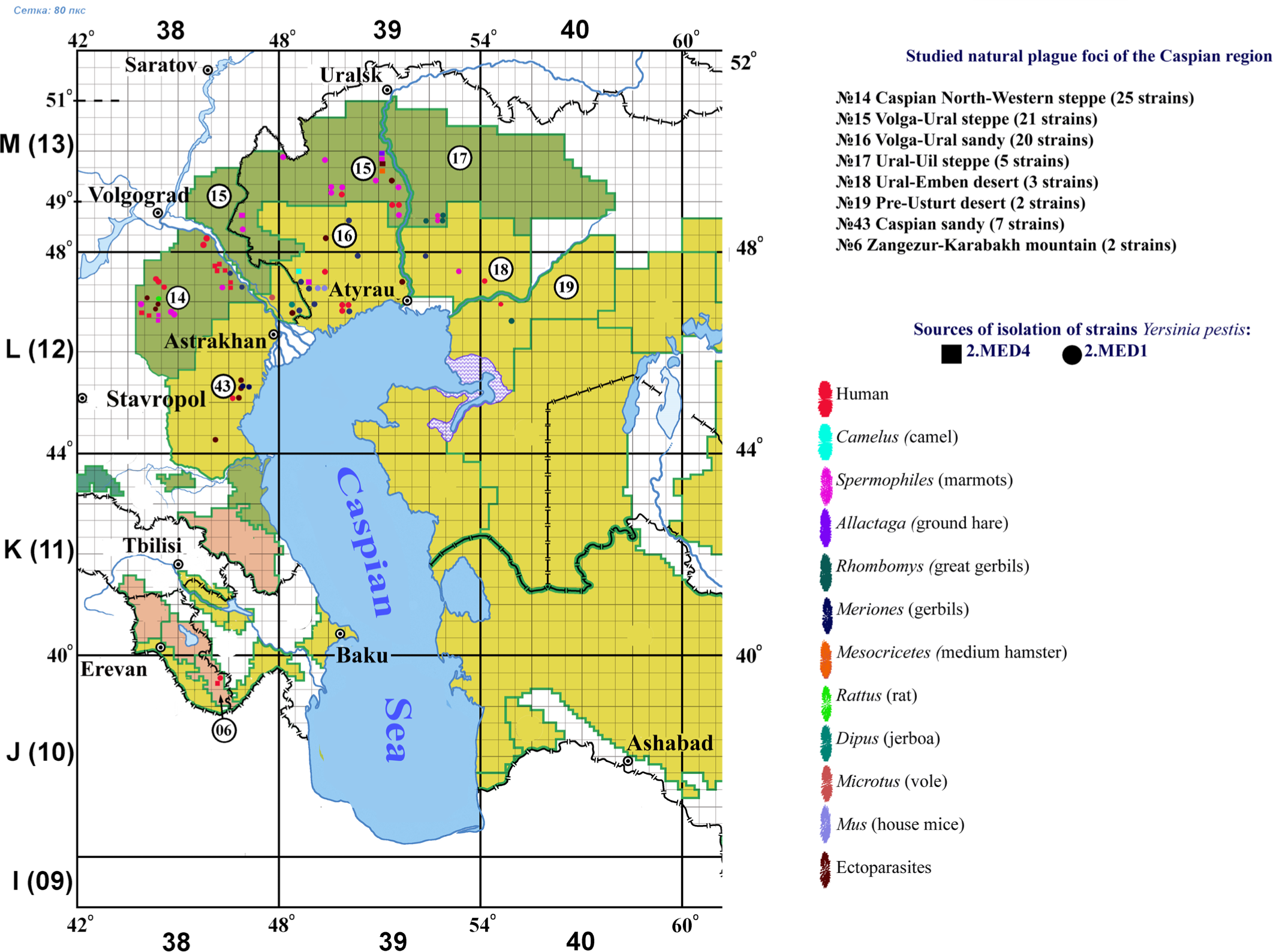
We studied 85 Y. pestis strains of the 2.MED1 and 2.MED4 phylogroups that were isolated from carriers, vectors, and humans in natural plague foci of the Caspian Sea region between 1912 and 2015. Eighteen strains (1917-1950) belonged to the 2.MED4 phylogroup. The geographical distribution and biological source of the studied strains are shown in Fig. 1.
The strains were cultivated on agar or LB broth (pH 7.2) at 28°C or 37°C for 24-48 h. A set of Hiss liquid media and commercial API 20E strips (Bio Merieux, France) were used to evaluate the enzymatic activity. To measure denitrifying activity, 108 CFU/ml of Y. pestis were inoculated in 5 ml of LB broth containing 0.1% potassium nitrate (KNO3) and then cultivated for 72 h at 28°C. The results were visualized by adding 0.5 ml of the Griess reagent: the medium that contains a strain exhibiting denitrifying activity should turn a crimson color. Medieval biovar strains are not capable of nitrate reduction [6].
The nutritional requirements of Y. pestis strains from the Caspian foci were determined according to the method proposed by Brubaker [7] with some modifications [8]. The results were visualized as a heatmap of a correlation matrix using the Seaborn v0.11.2 library.
The ability of the strains to form pigmented colonies was tested in three parallel inoculations on LB medium with Congo red dye. A suspension of daily culture in a concentration of 5x103 CFU/ml (0.1 ml) was spread over an agar plate and incubated at 28°C for 24 h, followed by 2-10 days at 4°C. The results were evaluated based on the presence of grown pigmented colonies; colorless colonies indicated the absence or inactivity of the chromosomal fragment that carries genes mediating colony pigmentation [6].
Higuchi & Smith medium (magnesium oxalate agar) was used to test whether the growth of plague microbe strains depends on the presence of calcium ions in the medium. An aliquot of 0.1 ml from each dilution of a daily culture suspension (1x103, 1x104, and 5x105 CFU/ml) was sown on a solid nutrient medium. The grown Ca2+-independent colonies were counted after 48 h of incubation at 37°C. Next, Ca2+-dependent colonies were counted after additional incubation at 28°C for 24 h. Y. pestis EV NIIEG was used as a control sample to set the pigmentation criterion and Ca2+ dependence [6]. Plasmid profiles of the Y. pestis strains were analyzed according to the method proposed by Kado and Liu [9]. Seventy-eight housekeeping and virulence genes of Y. pestis strains from the 2.MED4 and 2.MED1 phylogroups were compared by aligning gene regions to the reference sequence of Y. pestis CO92 (GenBank accession number NC_003143.1) using the MEGA X program, Exonerate [10], and the BLAST algorithm. We previously deposited the whole-genome sequences of the studied Y. pestis genomes in NCBI Gen-Bank [1, 3, 4].
RESULTS AND DISCUSSION
Biochemical properties, nutritional requirements, pigment sorption, and Ca2+ ion dependence
In this research project, we studied the main subspecies of the Y. pestis strains isolated from carriers, vectors, and humans in eight plague foci in the Caspian Sea region in 1912-2015: Caspian North-Western steppe (25 strains), Volga-Ural steppe (21 strains), Volga-Ural sandy desert (20 strains), Ural-Uil steppe (5 strains), Ural-Emben desert (3 strains), Pre-Usturt desert (2 strains), Caspian sandy region (7 strains), Zangezur-Karabakh mountain region (2 strains). The biochemical properties of all studied strains suggested a phenotype typical of the medieval biovar: denitrification−, fermentation of glycerol+, rhamnose−, and arabinose+. Using the API 20E test, we found that 18% of the studied Y. pestis strains had β-galactosidase activity: cleavage of the orthonitrophenyl-β-galactoside substrate with the release of ortho-nitrophenol and galactose. In addition, 5% of the strains were able to ferment amygdalin (glycoside, AMY). Such strains were isolated in the plague foci located in the Caspian North-Western steppe (11 strains), Volga-Ural steppe (4 strains), Volga-Ural sandy desert (2 strains), and Zangezur-Karabakh mountainous region (2 strains).
The analysis of 85 strains of Y. pestis from the Caspian Sea region showed that they had identical nutritional requirements for Phe, Met, and Thr regardless of the geographical area (Fig. 2A, B). Most strains isolated from the Caspian plague foci demonstrated Cys auxotrophy, with only 9% of the bacteria examined showing independence from Cys for their growth. Nine strains from the Caspian Sea region foci required Leu, which was consistent with the results of other studies [7]. Additionally, we found that the strains that belong to the first half of the 20th century required Arg (4 strains) and Ser (3 strains). We found no differences in nutritional requirements between the strains of the 2.MED1 and 2.MED4 phylogroups. The genetic factors underlying Arg, Ser, and Leu auxotrophy in several 2.MED1 and 2.MED4 strains, isolated during the first half of the 20th century, remain to be elucidated. The results of determining the nutritional needs of Y. pestis strains from the Caspian foci were visualized as a correlation matrix (heatmap) using the Matplotlibbased Seaborn v0.11.2 library in Python 3.9.0 (Fig. 2A, B). Calculation of the frequency of occurrence of nutritional needs for each amino acid was done for each studied focus (Fig. 2B).
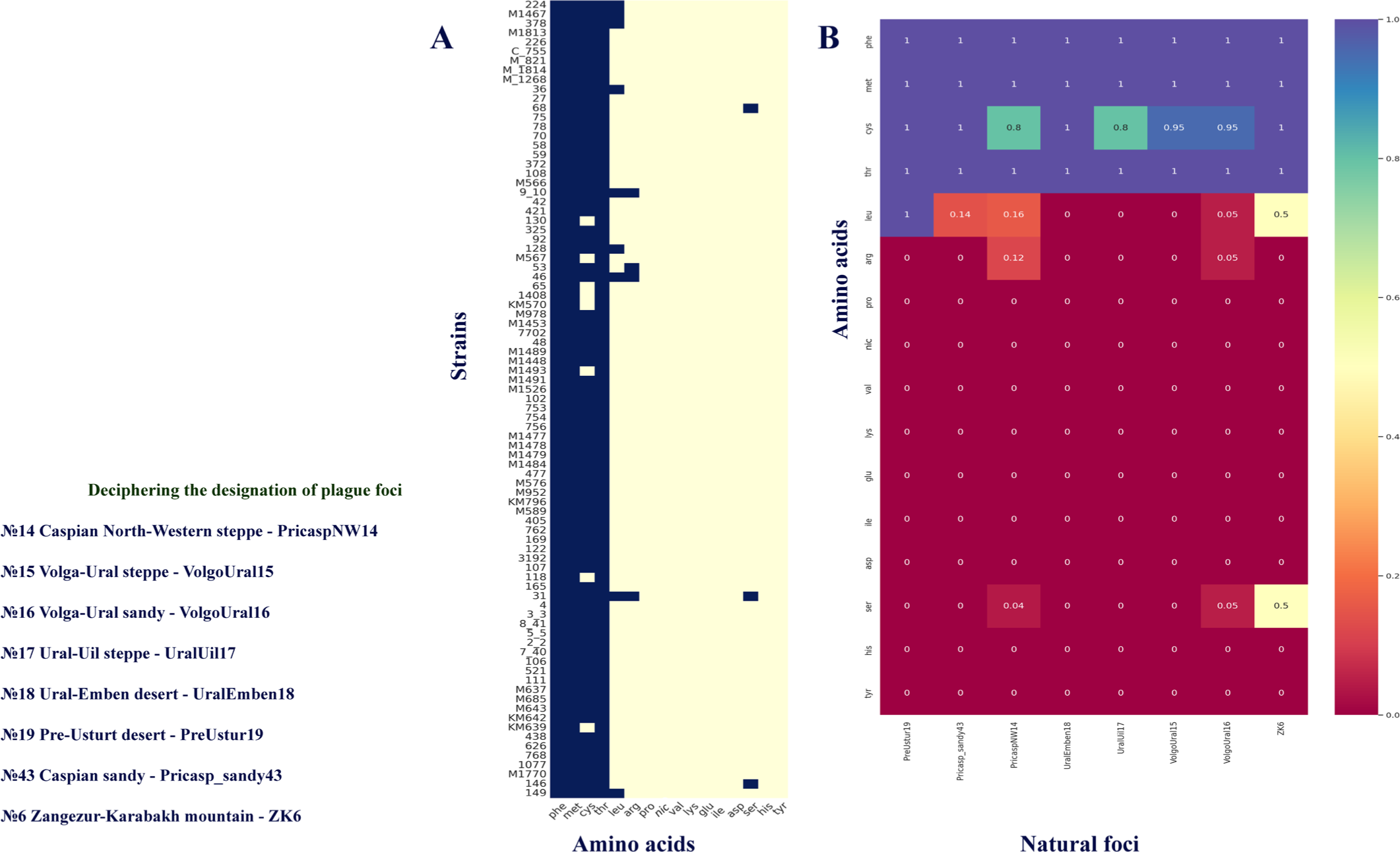
According to the results of analyses, the studied strains of the 2.MED4 and 2.MED1 phylogroups have a similar biochemical activity profile, which can indicate the phenotypic similarity of these populations due to adaptation to the same landscape, ecological, and faunal conditions in natural plague foci.
We examined 85 strains of Y. pestis for virulence-associated traits, such as pigment sorption and dependence of bacterial growth on the presence of Ca2+ ions in the medium. The loss of these traits leads to the attenuation of the plague microbe. An analysis of the ability of Y. pestis strains to absorb the Congo red pigment in LB medium showed that several strains lacked pigment sorption, as indicated by the formation of white colonies. Eight strains did not form pigmented colonies, while heterogeneity in this trait was noted in 24 strains (80-90% Pgm+; 60-70% Pgm−). Some of the studied strains from the early 20th century may have lost the 102-kb chromosome fragment from the pgm locus, which is responsible for this trait, due to its instability resulting from homologous recombination between two copies of flanking IS100 elements. The analysis of Y. pestis strains’ growth dependence on Ca2+ ions in Higuchi & Smith medium at 37°C showed that two strains of Y. pestis from the 2.MED1 phylogroup, which belonged to the first half of the 20th century, were Ca2+-independent. This observation indicates a potential loss of a virulence determinant, specifically the Ca2+-dependence plasmid pCD1.
Plasmid screening of Y. pestis from the 2.MED4 and 2.MED1 phylogroups
The plague pathogen carries three types of resident plasmids in addition to circular chromosomal DNA. The pCD1 plasmid that determines the Ca2+-dependence of bacteria is genus-specific. Y. pestis also carries species-specific plasmids pMT1 (encodes the Fraction 1 capsular antigen) and pPCP1 (encodes pesticin). These three plasmids contain virulence genes necessary for the infectious process or the transmission of plague through fleas. According to the results of screening, 16 strains from the first half of the 20th century lacked the pPCP1 plasmid. This was likely caused by long-term storage without lyophilization. The strains were isolated from different sources on the territory of the Caspian North-Western steppe (5 strains), Volga-Ural steppe (4 strains), and Volga-Ural sandy (7 strains) plague foci. Two Y. pestis 2.MED1 strains of the medieval biovar isolated in the first half of the 20th century lacked the Ca2+-dependence pCD1 plasmids, which was consistent with their ability to grow at 37°C on Higuchi & Smith medium without Ca2+ ions. The pCD1 plasmid encodes genes mediating virulence, and the plague microbe becomes avirulent if the plasmid is lost. The remaining studied strains of the medieval biovar of Y. pestis carried all three plasmids and were virulent.
Genetic determinants associated with virulence and housekeeping genes of Y. pestis 2.MED4 phylogroup strains in comparison with other strains of the medieval biovar
We compared 78 virulence and housekeeping genes of the Y. pestis strains from the 2.MED4 phylogroup with the corresponding genes of other phylogenetic groups within the medieval biovar, the full genome sequences of which we previously deposited in the NCBI GenBank database [1, 3, 4]. Genes encoded by the chromosome (43 genes) and three plasmids of the plague pathogen – pMT1 (3 genes), pCD1 (29 genes), and pPCP1 (3 genes) – were analyzed using the MEGA X, Exonerate, and BLAST. Chromosomal genes are involved in virulence manifestation, energy metabolism, membrane transport, and survival in nutrient deficiency conditions. These include the pigmentation genes and genes encoding adhesins (pH6 antigen and Ail protein), various enzymes, transporters, membrane proteins, and transcription factors. A synonymous substitution (111, C→T) was found in the chromosomal gene hemS that encodes the hemoglobin-degrading factor. This substitution is present in all phylogroups of the medieval biovar (Table 1).
| Gene location | Gene, SNP type, substitutions a | Encoded product | Phylogenetic affiliation |
|---|---|---|---|
| Chromosome |
hemS
111, C→T synonymous | Hemoglobin-degrading factor | Specific to 2.MED |
| pMT1 |
caf1M
29, G→A nonsynonymous 10, Gly→Glu | Periplasmic chaperone (as a part of the F1 capsular antigen) | Specific to 2.MED4 |
| pCD1 |
ssaJ
insertion of C (272) | Type III secretion system protein, YscD/HrpQ family | Specific to 2.MED1, 2.MED2, 2.MED3, 2.MED4 |
The location of a mutation is indicated relative to the position in the corresponding reference gene of the Y. pestis strain CO92 (AL590842.1).
A comparative analysis of the nucleotide sequences of caf1AM (encapsulation gene) and ymt (murine toxin gene) encoded by the pMT1 plasmid identified a nonsynonymous substitution (29, G→A, nsSNP: 10, Gly→Glu) in the nucleotide sequence of caf1M in 2.MED4 strains compared to 2.MED1 and other phylogenetic groups of Y. pestis. As a part of the encapsulation operon caf1, caf1M encodes the periplasmic chaperone protein. The substitution corresponds to the 10th amino acid position in the protein sequence and does not seem to disrupt the functional activity of the protein, which was confirmed by an immunochromatographic test for the presence of the F1 capsular antigen (State Scientific Center for Applied Microbiology and Biotechnology, Obolensk, Russia).
We analyzed 29 genes located on the pCD1 plasmid, encoding the Yop virulon (yopEHJKMT, ypkA), components of the type III secretion system (yscBCDEFGK-LOPQRSTUYVX, ssaJ, lcrRO), and V antigen (lcrV). The insertion of cytosine (C) at position 272 was found in the ssaJ gene, which encodes a protein of the type III secretion system that belongs to the YscD/HrpQ family. This insertion resulted in a frameshift mutation and gene function impairment. Since the Ysc system consists of 25 proteins, the function of the ssaJ encoded protein could likely be taken over by other proteins of this complex. This mutation is specific to the 2.MED1, 2.MED2, 2.MED3, 2.MED4 branches of the medieval biovar, but not for 2.MED0 (Table 1). No variability in the pPCP1 plasmid genes encoding the plasminogen activator (pla), pesticin (pst), and pesticin immunity protein (pim) was found between the strains of the medieval biovar. The studied virulence genes and housekeeping genes showed a high degree of conservation. This could be explained by the importance of their products for Y. pestis pathogenicity, survival, and viability, as well as by the evolutionary youth of the medieval biovar of the plague microbe.
To summarize, we analyzed the phenotypic and genotypic characteristics of Y. pestis strains of the medieval biovar from the 2.MED1 and 2.MED4 phylogroups – the etiological agents of plague outbreaks in the Caspian Sea region in the early 20th century. Our comprehensive study, which examined the biochemical properties and nutritional requirements of 85 Y. pestis strains from the Caspian plaque foci, revealed no significant differences between the 2.MED4 and 2.MED1 strains at the phenotypic level.
We found that all studied 2.MED4 and 2.MED1 strains from these natural foci had a nutritional requirement for Met, Phe, and Thr. In addition, most of the strains required Cys: only 9% of the strains did not exhibit this requirement. We showed that some of the Y. pestis strains from the Caspian region that belonged to the first half of the 20th century also required Arg (4 strains) or Ser (3 strains). Some strains from the first half of the 20th century lacked plasmids pPCP1 (pesticin production) and pMT1 (Ca2+-dependence). It was also found that several strains lacked the functional chromosomal region of pigmentation, as suggested by their inability to form pigmented colonies. The analysis of 78 essential genes and virulence genes provided new data on the genetic variability of Y. pestis. Medieval biovar strains exhibited variability of the hemS gene encoding the hemoglobin-degrading factor. In the 2.MED4 phylogroup of the plague microbe, we identified a mutation in the caf1M gene of the periplasmic chaperone, which represents a part of the operon regulating the synthesis of the virulence factor – capsule protein F1. Lastly, the 2.MED1, 2.MED2, 2.MED3, and 2.MED4 phylogroups of the medieval biovar showed variability in the ssaJ gene, which encodes a protein of the type III secretion system from the YscD/HrpQ family.
The absence of significant phenotypic heterogeneity in differential biochemical properties between the 2.MED4 and 2.MED1 phylogroups can be attributed to their relatively short period of evolutionary divergence and the similar habitat conditions in the natural ecosystems of the Caspian region. Based on our findings reported in this and other publications, along with data from epizootological monitoring, we can propose a sympatric scenario for the formation of these populations of the medieval Y. pestis biovar [3-5]. The implications of the discovered single nucleotide substitutions in the virulence genes and essential genes, which have caused the divergence of these phylogroups, are to be explored. Further studies on the phenotypic and genotypic features of the medieval biovar strains of Y. pestis could help determine the genetic basis of auxotrophy and the specifics of the biochemical profile of this biovar. These findings can be utilized in laboratory diagnostics of the plague microbe, as well as for the characterization of the genetic relatedness of these highly virulent populations of Y. pestis in the modern period.

