Introduction
Septoria tritici blotch caused by Zymoseptoria tritici (Dersm.) Quaedvlieg & Crous, is the most destructive disease of winter wheat in Ireland, annually requiring intensive fungicide applications during the growing season to prevent significant yield losses. The development and spread of fungicide resistance in Z. tritici is a serious threat to the viability of winter wheat production in Western Europe (Jess et al., 2014; Blake et al., 2018; Jørgensen et al., 2021). Unfortunately, Z. tritici strains, completely or partially resistant to the major fungicide modes of action available for use on wheat (e.g. azoles, quinone outside inhibitors [QoIs] and succinate dehydrogenase inhibitors [SDHIs]), are now present in Z. tritici populations throughout the region (Cools & Fraaije, 2013; Dooley et al., 2016a; Huf et al., 2018; Hellin et al., 2021). Depending on the frequencies of resistances in these populations, the efficacy of the different fungicide classes can be severely impacted (Kildea et al., 2010; Blake et al., 2018; Jørgensen et al., 2021). To date, resistance is predominantly conferred through the development of changes in the different fungicide target sites, either preventing or reducing the capacity of the fungicide to bind to its target (Fisher & Meunier, 2008; Cools et al., 2011; Fraaije et al., 2012). Additional resistance mechanisms, including overexpression of target sites or efflux pumps, have also been identified in European Z. tritici populations (Cools et al., 2012; Omrane et al., 2017), although their impact on the field efficacy of those specific fungicide classes affected remains largely unknown.
A total of 25 alterations in 14α-demethylase (CYP51), the target site of the azole fungicides, have been detected in European Z. tritici strains, with multiple combinations of these alterations now present in individual strains (Cools & Fraaije, 2013; Huf et al., 2018). Their development and accumulation/combination has occurred in a stepwise manner, most likely in response to the constraints imposed by specific alterations on the functioning of the protein and compensatory effects brought by others (Cools et al., 2011). Understanding the potential interactions between the different alterations and their impacts on azole sensitivity is important to ensure the careful development of fungicide programmes that are needed to provide adequate disease control, whilst limiting the selection of resistant strains. For instance, as highlighted by Fraaije et al. (2007) and later by Dooley et al. (2016b), the potential exists to utilise both the diversity in azoles and their different impacts on local Z. tritici populations for both disease control and resistance management.
In recent years, although the ability to monitor CYP51 haplotypes has greatly increased through the development of long-read sequencing (Samils et al., 2021), due to both the high cost and limited accessibility, such methods are unlikely to be readily available to routinely monitor changes in azole sensitivity. Hellin et al. (2021) recently demonstrated the flexibility and continued value in quantitative polymerase chain reaction (qPCR) assays targeting specific alterations, with clear differences in specific alterations relating to both azole and SDHI resistance observed across Europe, reflecting the differences in disease pressures and fungicide usage. qPCR assays are limited in their capacity to detect a small number of alterations in a single assay (dependent on assay design and technology available). The choice of specific alterations associated with azole resistance to target in these assays is vital to ensure the relevance of the findings.
Using collections of Z. tritici established in Ireland over the past decade and capturing the diversity of sensitivity/CYP51 haplotypes present in contemporary European populations, we have applied both conditional inference trees and random forest to identify which specific alterations in Z. tritici CYP51 and their relationships confer the greatest impact on azole sensitivity (epoxiconazole [EPZ] and metconazole [MTZ]). The relationships identified confirm the importance of the key alterations both I381V and S524T to both azoles, supporting the findings of Cools et al. (2011), and validate the selection of S524T as a key marker for azole resistance in contemporary populations by Hellin et al. (2021).
Materials and methods
Zymoseptoria tritici isolate collections and azole sensitivity
The isolates examined in the present study were selected from three previously published collections. These include a subsample of: the 2012 collection (n = 506) established and characterised by Dooley et al. (2016b), a 2015 collection (n = 78) described by Kildea et al. (2019), and the 2017 collection described by Samils et al. (2021). They are representative of the wider Irish Z. tritici population over the past decade and each was retrieved from winter wheat cultivars typically grown during this period. In the case of both the 2012 and 2015 collections, the isolates originated from fungicide-treated wheat plots or crops (i.e. trial plots in 2012 and commercial crops in 2015). The 2017 collection was established from infected leaves sampled prior to the application of fungicides. All isolates were stored as spore suspensions in 30% glycerol at −80°C and revived by spotting 30 μL of the suspension on potato glucose agar and subsequently incubated at 18°C in darkness. The sensitivity of each isolate to the azoles EPZ and MTZ was previously determined by Dooley et al. (2016b), Kildea et al. (2019) and Samils et al. (2021). Cross-resistance to both chemicals was determined using Pearson’s correlation.
CYP51 sequence analysis and upstream inserts detection
For both the 2015 and 2017 collections, DNA extraction and CYP51 sequence analysis were as described by Kildea et al. (2019) and Samils et al. (2021), respectively. For the 2012 collection, DNA was extracted following freeze-drying of yeasty spores collected after 4 d growth on poly-γ-glutamic acid potato glucose agar (PGA), using a NucleoSpin® 96 Plant II extraction kit (Macherey-Nagel, Düren, Germany). For each collection, PCR amplification and sequencing of the entire CYP51 gene was performed as described by Kildea et al. (2019). Sequence assembly and alignment was performed in CLC workbench with each individually assessed using Bioedit. Amino acid haplotypes were identified in accordance with the nomenclature devised by Huf et al. (2018), with novel haplotypes assigned the next in sequence. Analysis of the different isolates for insertions in the promoter was as described by Cools et al. (2012) and Kildea et al. (2019).
Impact of CYP51 alterations on epoxiconazole and metconazole sensitivity
Conditional inference trees were used to study and visualise the impact of amino acid changes in CYP51 on EPZ and MTZ sensitivity. To limit potential interference from strains with CYP51 overexpression, strains identified with the 120-bp CYP51 promoter insert were excluded from the analysis. Construction of the conditional inference trees was carried out using the ctree function from the R “party” package (Hothorn et al., 2010) with the CYP51 amino acid alterations as predictors and either EPZ or MTZ as the response variable. The distribution of the test statistic was computed using Bonferroni (testtype = “Bonferroni”), the value of the test statistic to exceed before implementing a split was set to 1 – (P-value of 0.05) (mincriterion = 0.95), the minimum sum of weights in a node before considering a split was set to 40 (minsplit = 40) and the minimum sum of weights in a terminal node was set to 20 (minbucket = 20). The tree was plotted in R. Measures of variable importance were also calculated using the R package RandomForest (Liaw & Wiener, 2002). Random forest regression (Breiman, 2001) was run with the number of variables randomly sampled at each split set to 6 (mtry = 6), the number of trees to grow set to 500 (ntree = 500) and the importance of predictors assessed (importance = TRUE). The mean decrease in accuracy was plotted for each variable.
Results and discussion
A wide range in sensitivity was observed in the collections to both azoles (Figure 1). For EPZ, a resistance factor (RF: ratio of most to least sensitive) of 2,486 was detected amongst the collections. The mean sensitivity to EPZ decreased from a half maximal effective concentration (EC50) value of 0.73 mg/L in the 2012 collection to 2.02 mg/L in 2015 and 1.86 mg/L in 2017. Although care must be taken in the interpretation of these differences as each collection was collected at different sampling time points and hence fungicide treatments, it is reflective of the wider decrease in the sensitivity of the Irish Z. tritici population as previously reported by Kildea et al. (2019).
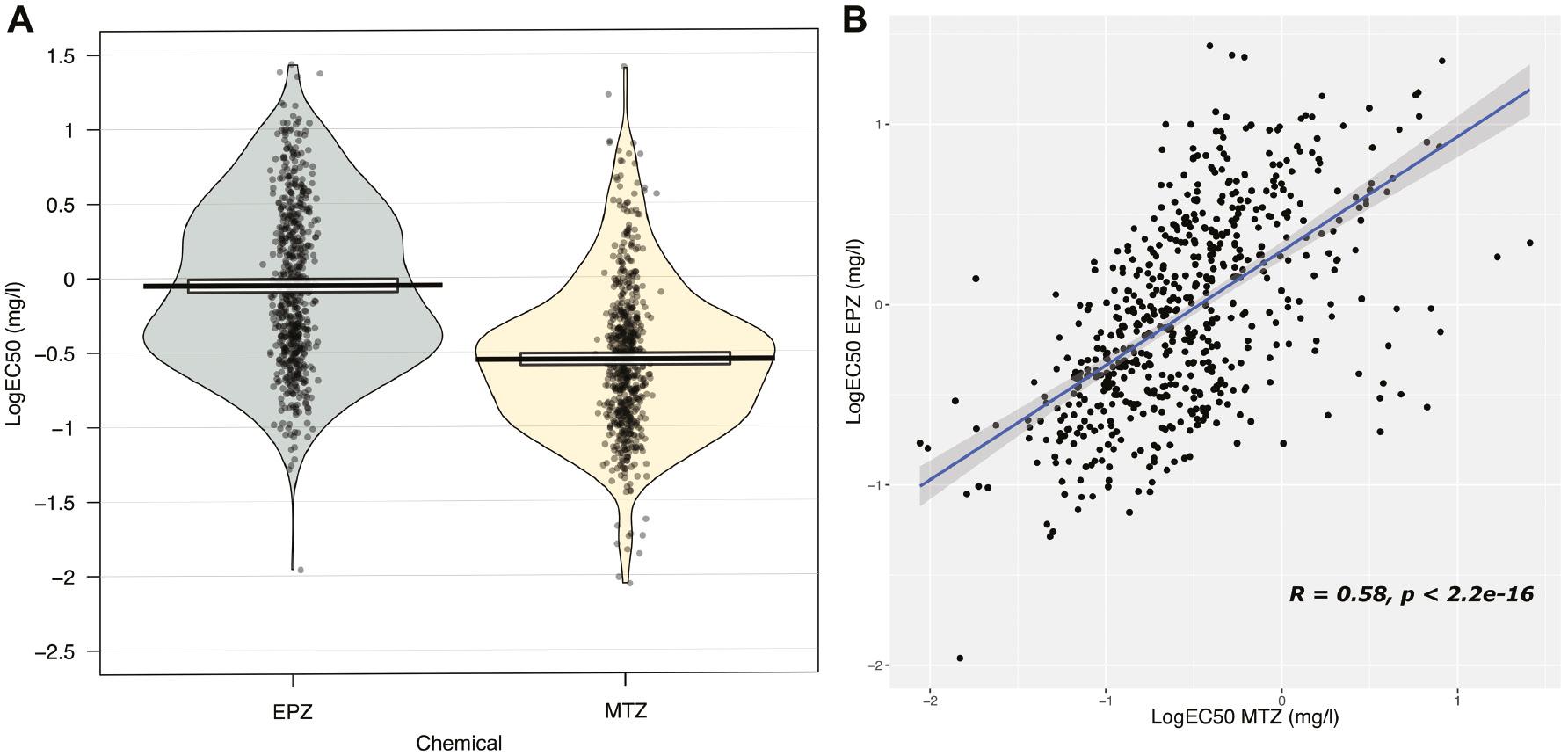
(A) Distribution of sensitivity of the Zymoseptoria tritici isolates included in the study to epoxiconazole (EPZ) and metconazole (MTZ). Data are presented using a pirateplot with density plot, central tendency, and 95% confidence interval. (B) Cross-resistance between both fungicides. Sensitivity is represented as LogEC50 (mg/L).
A similar range in sensitivity was observed to MTZ, with an overall RF in the collection to MTZ of 2,941. The mean sensitivity to MTZ also decreased, from an EC50 value of 0.24 mg/L in 2012, to 0.64 mg/L in 2015 and 0.60 mg/L in 2017. As per the EPZ sensitivity, caution must be observed in the interpretation of these differences, but again they are representative of the change that has occurred in the Irish population. The distribution of sensitivity of the collections to both chemicals was unimodal, with both slightly skewed in opposing directions (Figure 1). A significant Pearson’s correlation coefficient (R = 0.58; P < 0.001) was detected in the sensitivity of the entire collection to EPZ and MTZ (Figure 1B), suggesting common mechanism(s) of resistance towards each.
Amongst the isolates, 23 CYP51 amino acid alterations at 17 different positions were detected (Figure 2A). All alterations detected have previously been reported in European populations (Figure 2A) (Cools & Fraaije, 2013; Huf et al., 2018; Huf et al., 2020). The most common alteration L50S was detected in 99.4% of the isolates. The alterations which have commonly been used as markers for azole resistance V136A, I381V and S524T (Jørgensen et al., 2021) were found in 65.5%, 79.2% and 54.0% of the isolates, respectively. The 120-bp insert previously identified as conferring increased CYP51 expression by Cools et al. (2012) was identified in 7.4% of the isolates.
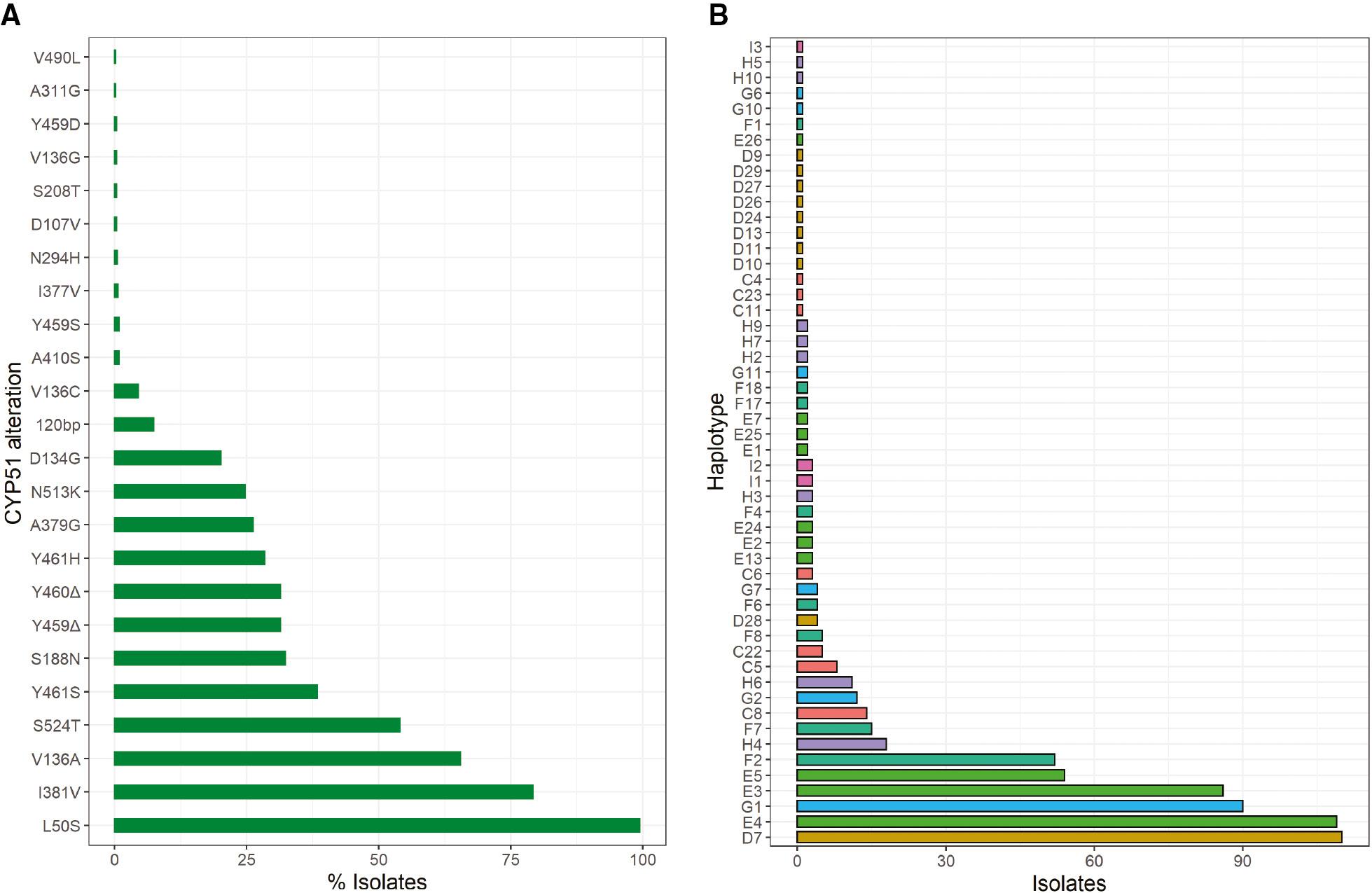
Frequency of (A) specific CYP51 amino acid alterations and (B) CYP51 haplotypes based on nomenclature devised by Huf et al. (2018) in the Zymoseptoria tritici collections analysed.
Comparable to previous European monitoring studies by Huf et al. (2018), alterations were detected in multiple combinations. In total, 52 different combinations were identified in the collections (Figure 2B). In most instances the different combinations were successfully assigned a haplotype based on the nomenclature as developed by Huf et al. (2018) (Figure 2B). No wild-type haplotypes were found and no isolates with either a single alteration or only two alterations were present in the collection. At the other extreme seven isolates had nine alterations. The most common haplotypes detected included D7 (L50S, V136A, Y461S, S524T), E4 (L50S, D134G, V136A, I381V, Y461H), G1 (L50S, S188N, A379G, I381V, Δ459/460, N513K) and E3 (L50S, V136A, I381V, Y461S, S524T), and combined accounted for almost 60% of all isolates. The haplotypes E4, G1 and E3 were also the most commonly identified throughout Europe in 2016 by Huf et al. (2018). The 120-bp CYP51 promoter insert, which confers CYP51 overexpression, was only identified in isolates with the F2 (L50S, S188N, I381V, Δ459/460, N513K) haplotype.
To understand and visualise how the different amino acid alterations potentially influence sensitivity to EPZ and MTZ, conditional inference trees were generated. To avoid overrepresentation of alterations that may have arisen due to the presence of strains with CYP51 overexpression, strains with the 120-bp CYP51 promoter insert which has been shown to confer CYP51 overexpression (Cools et al., 2012; Kildea et al., 2019) were excluded from the analysis. For both EPZ and MTZ, inference trees demonstrated that a combination of the amino acids V136A/C, I381V and S524T led to a decrease in sensitivity (Figure 3). Random forest was subsequently used to calculate variable importance measures and determine the proportion of variance in sensitivity that could be accounted for by these alterations. For EPZ 64.15% of the variance in sensitivity could be explained by the identified CYP51 alterations. This decreased to 40.25% for MTZ sensitivity. For both chemicals, the importance of both S524T and I381V was highlighted, with both alterations displaying the greatest importance for the respective model accuracy (Figures 3B and 4B). For both chemicals, it should be noted that a significant proportion of the variance in sensitivity was not explained by the various CYP51 alterations detected. Omrane et al. (2017) previously demonstrated the role of MFS1 in reducing azole sensitivity in Z. tritici, and this or alternative efflux pumps may play also a role in the isolate collections examined.
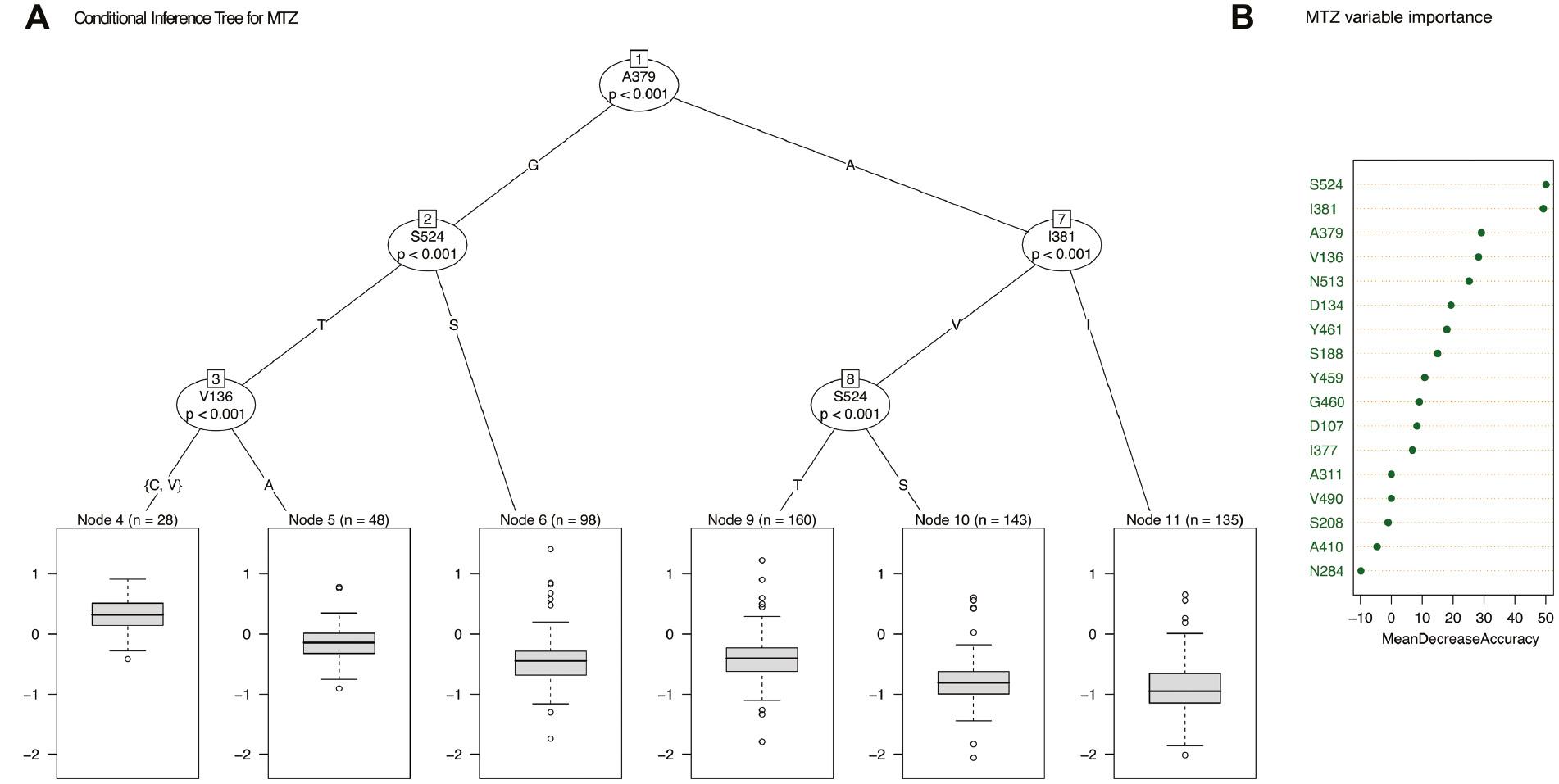
(A) Conditional inference tree with the metconazole (MTZ) LogEC50 (mg/L) as response variable and amino acid changes as predictors. The box plots show MTZ LogEC50 (mg/L) for samples within each node (n = sample number), and amino acid substitutions are shown on tree. (B) Variable importance plot showing how model accuracy decreases as we drop variables (generated using RandomForest).
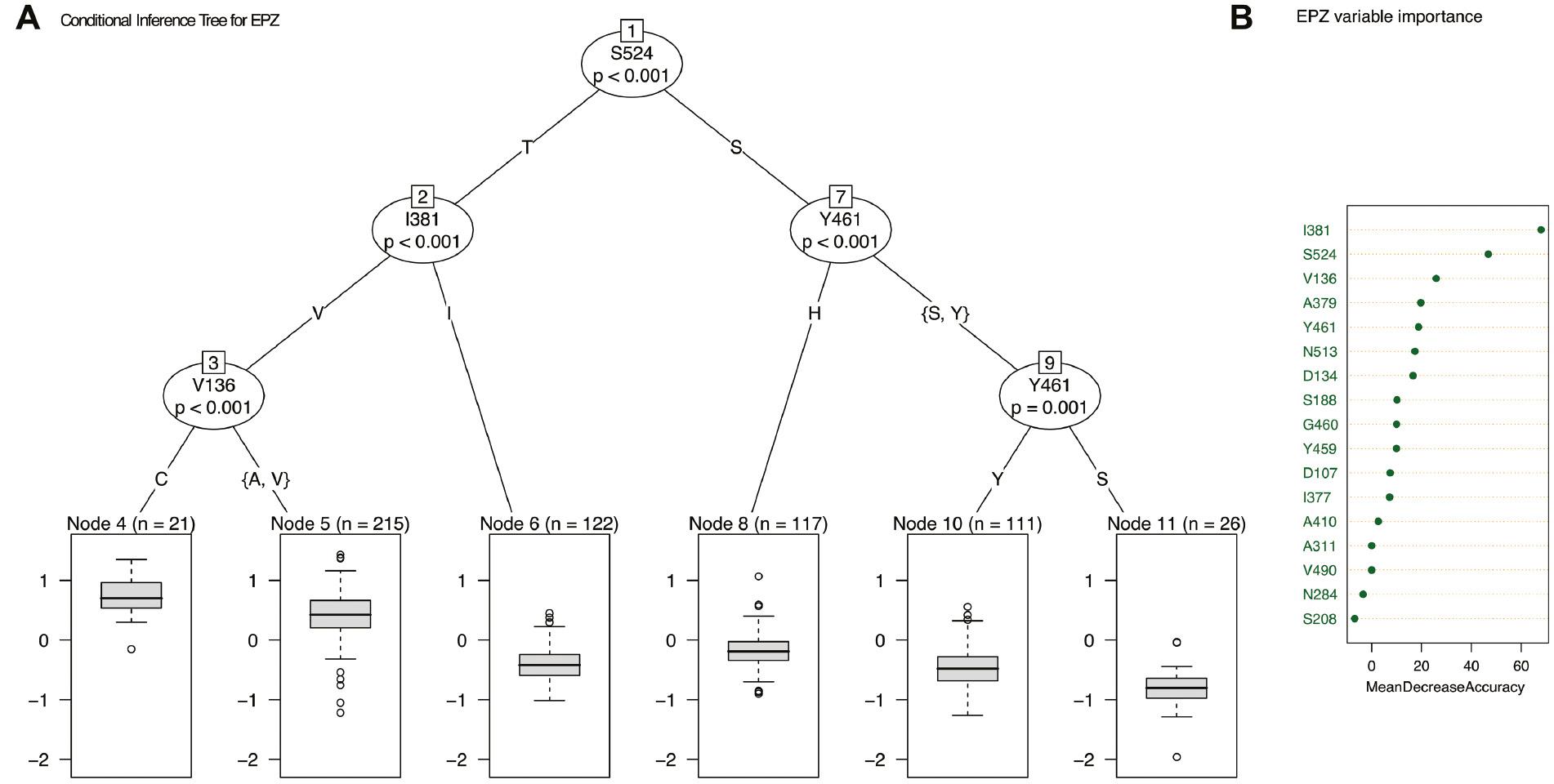
(A) Conditional inference tree with the epoxiconazole (EPZ) LogEC50 (mg/L) as response variable and amino acid changes as predictors. The box plots show EPZ LogEC50 (mg/L) for samples within each node (n = sample number), and amino acid substitutions are shown on tree. (B) Variable importance plot showing how model accuracy decreases as we drop variables (generated using RandomForest).
The I381V alteration was first described by Fraaije et al. (2007), in which the authors suggested that the alteration negatively impacted azole sensitivity, in particular tebuconazole. Through heterologous expression of mutated ZtCYP51 in yeast, the potential role of I381V was further explored by Cools et al. (2011). The authors found that in addition to negatively impacting azole sensitivity the alteration alone was deleterious to 14α-demethylase, demonstrating that the accumulation of alterations not only accommodated further reductions in azole sensitivity, but was also required to facilitate the emergence of specific alterations. Since then, I381V has increased throughout European Z. tritici populations, with most populations now dominated by strains with the alteration (Jørgensen et al., 2021). Using the same heterologous expression system, Cools et al. (2011) also demonstrated the impact of S524T on azole sensitivity, with the alteration leading to significant reductions in azole sensitivity in numerous combinations with other alterations. Using molecular modelling, Cools et al. (2011) also suggested that the impact on sensitivity conferred by these alterations is in part due to an increase in the azole binding cavity. Again, similar to I381V, the prevalence of S524T in European Z. tritici populations continues to increase, although a gradient in frequency from high in Western Europe to low in Eastern Europe is still observed (Hellin et al., 2021; Jørgensen et al., 2021).
For both EPZ and MTZ, the combination of alterations with the greatest impact on sensitivity to both chemicals as identified by the conditional inference trees (Figure 3) included alterations at position 136, in particular the alteration V136C, in combination with I381V and S524T. The haplotype H6 which has this combination of alterations, in addition to L50S, A379G and Δ459/460, was also amongst the least sensitive to EPZ as identified by Huf et al. (2018). Although position 136 was amongst the highest predictors of reduced sensitivity for both EPZ and MTZ in the random forest, it was ranked lower in importance when compared to either I381V or S524T, further highlighting the cumulative impact of key alterations. It also highlights potential difficulties for future molecular monitoring of azole resistance in Z. tritici, with the potential needed to monitor haplotype frequencies instead of specific alterations. This will be greatly facilitated with further application of long-read sequencing for resistance detection such as that developed by Samils et al. (2021) or variations of the potentially more accessible MARPLE point-of-care approach developed for Puccinia striiformis f. sp. tritici and based on long-read sequencing Oxford Nanopore sequence technology (Radhakrishnan et al., 2019).

