INTRODUCTION
The direct targets of antibiotics have long been a primary focus of research. Resistance mechanisms of quinolone antibiotics are well-documented and often involve specific alterations in bacterial enzymes. Bacterial DNA gyrase, a tetramer comprising two gyrA and two gyrB subunits, develops high-level quinolone resistance, primarily through genetic alterations [1,2]. Resistance arises through point mutations in specific areas of gyrA and gyrB, known as quinolone resistance-determining regions [3–6]. Novobiocin (NV), an antibiotic belonging to the coumarin class, targets DNA gyrase and shows efficacy against resistant staphylococcal infections [7]. Resistance to NV results primarily from mutations in the gyrB gene, particularly near the gyrB ATP-binding pocket [8–10]. However, the mechanisms of NV resistance are complex and remain incompletely understood, particularly regarding the antibiotic’s effects on specific metabolic pathways in S. aureus. Understanding the effects of these antibiotics on the fundamental metabolic processes of S. aureus might enhance therapeutic strategies, provide insights into the downstream effects of DNA gyrase mutations, and support the rational design of effective treatments.
Mutations in the gyrB gene are frequently identified in ciprofloxacin (CIP) resistant S. aureus strains [11]. The mutation rate is approximately 23.5%, in agreement with the rates reported for both methicillin-resistant S. aureus (MRSA) and methicillin-sensitive S. aureus (20–25%) [12]. Although most reports of gyrB mutations are associated with CIP, exploring the biological functions of gyrB mutations induced by NV is crucial. This study used adaptive laboratory evolution (ALE) to develop NV-resistant S. aureus strains with alone gyrB mutations and used metabolomic analyses to compare these evolved strains with their ancestral counterparts. After acquiring NV resistance mutations, S. aureus was found to undergo extensive compensatory metabolic reprogramming, independently of the drug’s presence. This reprogramming affected purine and pyrimidine metabolism, as well as amino acid biosynthesis, thus providing new insights into the mechanisms of NV resistance in S. aureus.
MATERIALS AND METHODS
Bacterial strains, media, and reagents
The S. aureus strain ATCC25923 was cultured under BSL-2 conditions at 37°C. For bacterial monoclonal isolation, Baird-Parker agar supplemented with 5% potassium tellurite egg-yolk reagent was used. Routine cultivation was performed with tryptic soy broth (TSB) and tryptic soy agar (TSA). Antibiotics, including ampicillin (A430258), gentamicin (A428430), vancomycin (A414413), erythromycin (A429932), rifampicin (A430385), minocycline (A429552), cefalexin (A600280), kanamycin (A430277), streptomycin (A610494), tetracycline (A430165), chloramphenicol (A429048), moxifloxacin (A430708), levofloxacin (A417584), gatifloxacin (A428581), CIP (A427057), norfloxacin (A428525), ofloxacin (A430176), and novobiocin (A430130), were purchased from Shanghai Sangon Bioengineering Co., Ltd. Fusidic acid (R006290) and linezolid (R130584) were purchased from Shanghai Yien Chemical Co., Ltd. All compounds were filtered through 0.22-μm filters under sterile conditions before use.
Adaptive laboratory evolution of S. aureus with novobiocin
The ALE experiment was conducted according to established protocols with slight modifications [13,14]. In the first phase, TSB plates containing NV at concentrations ranging from 1× to 1,024× the minimum inhibitory concentration (MIC) (0.08–81.92 μg/mL) were prepared. An initially high amount of inoculum of S. aureus (~108 CFU/mL) (0.02 mL) was evenly spread onto 1× MIC plates and incubated at 37°C until visible colonies appeared. In the second phase, three clonal strains were selected from the 1× MIC plates and sub-cultured into fresh TSB liquid medium containing NV at the corresponding MIC concentration. The cultures were incubated at 37°C with shaking until turbidity was observed. At the stationary phase, the bacterial suspensions were centrifuged at 6,000 × g for 10 minutes, washed twice with PBS, and re-inoculated onto TSB plates containing increasing concentrations of NV. This process was repeated for ten passages while the antibiotic concentration was gradually increased to a maximum of 1,024× MIC (81.92 μg/mL). Each generation cycle required approximately 3 days and was conducted for at least ten passages. To assess resistance stability, we cultured the evolved strains in antibiotic-free medium for ten generations, and each generation was subsequently subjected to drug susceptibility testing. All experiments were performed in triplicate to ensure reproducibility.
Growth curve and minimum inhibitory concentration testing
Bacteria were cultured in TSB with shaking at 200 rpm under 37°C or on TSA unless otherwise specified. Permanent stock cultures were maintained in TSB containing 20% glycerol at −80°C, whereas working stock cultures on TSA were stored at 4°C. Overnight cultures were initiated from single colonies and incubated at 37°C with shaking at 200 rpm. Growth curves were initiated by inoculation of broth cultures with an overnight culture to achieve an initial OD600 nm of 0.04, as monitored over time in triplicate samples at 37°C and 200 rpm.
The MIC values for all isolates were determined with the broth microdilution method, in accordance with the Clinical and Laboratory Standards Institute guidelines. Serial two-fold dilutions of NV were prepared in TSB for susceptibility testing. After inoculation, the plates were incubated at 37°C for 8 hours; subsequently, 20 μL 0.01% (w/v) resazurin solution was added to each well. Overnight incubation allowed for bacterial growth to be observed according to a color change from blue to pink. The MIC was defined as the lowest concentration of NV that visibly inhibited bacterial growth. Because the Clinical and Laboratory Standards Institute guidelines do not provide breakpoint standards for S. aureus susceptibility to NV, we referred to the evaluation criteria established by Sweeney et al. [15], defining resistance as R ≥8 μg/mL. For drug cross-studies, interactions between drug pairs were evaluated according to the MICs, to assess potential synergistic, additive, or antagonistic effects.
Genomic extraction and whole-genome sequencing
S. aureus cells were cultured on TSB medium at 37°C for 18 h, then used for genomic DNA extraction. The genomic DNA was extracted with the cetyltrimethylammonium bromide method, with minor modifications [16]. Whole genome sequencing was performed by Shanghai Personal Biotechnology Co., Ltd. (China) as previously described [17].
Survival curves
Mid-exponential phase cultures of S. aureus were diluted in TSB medium, grown at 37°C, and treated with H2O2, N2H4, and SDS at the indicated concentrations. After the indicated time, cell survival was estimated with colony formation assays on TSB agar. All experiments were repeated at least three times.
Ethidium bromide and Nile red uptake assays
Mid-exponential phase cultures of the S. aureus strains were centrifuged and washed with PBS containing 0.05% Tween 80. The bacterial cell concentration was adjusted to an absorbance of ~0.4 at 600 nm. The bacterial cells were incubated in PBS with 2 μg/mL ethidium bromide and 2 μM Nile red in a black 96-well plate. Dye accumulation was measured with an excitation wavelength of 545 nm and emission wavelength of 600 nm for ethidium bromide, and an excitation wavelength of 540 nm and emission wavelength of 630 nm for Nile Red.
Metabolic activity
Metabolomic analysis of S. aureus was performed according to MRSA protocols [18]. To minimize cell death and lysis while capturing significant lethality at later stages, we used S. aureus with an OD600 value of 0.5 and exposed the bacteria to NV at 10× MIC for 60 minutes, according to the bactericidal curve obtained from our NV metabolism collection. This approach allowed us to observe the initial metabolic responses before cell death and compare them with those during the lethal phase [19]. After centrifugation to collect the bacteria, the samples were frozen at −80°C with liquid nitrogen, then thawed at 4°C. Each cell sample was subsequently resuspended in 300 μL pretreatment reagent containing a specific ratio of methanol, acetonitrile, and water (2:2:1 methanol:acetonitrile:water), and mixed with ceramic grinding beads. The mixture was then ground at 4°C 8,000 rpm for 5 minutes. The samples were centrifuged at 12,000 rpm at 4°C for 20 minutes, and 260 μL supernatant was transferred to a new centrifuge tube. The supernatant was concentrated to dryness with a centrifugal concentrator under vacuum at 45°C for 1 hour. Subsequently, 50 μL methanol/water resuspension reagent (3:7 methanol:water) was used to resuspend the sample, which was vortexed and centrifuged again at 12,000 rpm at 4°C for 20 minutes. Finally, 40 μL supernatant was placed in a sample vial (with an insert) for testing. Additionally, 5 μL was aspirated from each test sample, mixed in equal volumes, and vortexed to prepare the quality control sample. Protein concentrations in the supernatant were measured with a Bradford assay kit (Beyotime, China).
Metabolites were separated with a UHPLC system (Thermo fisher USA) coupled to a Q Exactive mass spectrometer (Thermo Fisher, USA). The QE MS instrument was used with the following specific conditions. The ion source was HESI, operating in both positive and negative ion modes. The scan mode was DDA, which entailed one full scan followed by ten MS/MS scans. The collision energy was set at 15 and 30 eV to fragment the ions, with high-purity nitrogen (99.999%) used as the collision-induced dissociation gas. The full scan range was set from 67 to 1,000 m/z, with a resolution of 70,000, an automatic gain control of 1e6, and an ion residence time of 100 ms. For the second-order mass spectrometry (dd-MS/MS), the resolution was 17,500, the automatic gain control was 5e5, and the ion residence time was 50 ms. The spray voltage was set to 3.2 kV in positive ion mode and 2.8 kV in negative ion mode. The ion source temperature was adjusted to 320°C, and the s-lens voltage was set to 50 V. The chromatographic column was an ACQUITY UPLC HSS T3 (100 × 2.1 mm, 1.7 μm, Waters), with the column temperature maintained at 40°C. The mobile phase consisted of water containing 0.1% formic acid (A) and acetonitrile containing 0.1% formic acid (B). The gradient elution program transitioned from A/B 99/1 to 0/100 within 12 minutes and was held at 0/100 until 13 minutes. The flow rate was 0.4 mL/min, and the injection volume was 1 μL. Data were collected in Xcalibur 3.0 software (Thermo fisher, USA). Peak extraction, alignment, and normalization were performed in Progenesis QI software (Waters, UK) for raw data. Metabolites were identified according to their accurate mass (with a mass error less than 5 ppm), MS/MS data, and comparison with the Human Metabolome Database (HMDB) conducted with TraceFinder (Thermo Fisher, USA). Differentially expressed metabolites (DEMs) were identified according to log fold change thresholds of ≥0.58 or ≤−0.58, with a p-value ≤ 0.05. Statistical analysis was performed with MetaboAnalyst 5.0 (http://www.metaboanalyst.ca/MetaboAnalyst/).
Statistical analysis
All data are presented as mean ± SD from at least three independent experiments and were analyzed with GraphPad Prism 8.0.1 software. Statistical significance was defined as p < 0.05 (*), p < 0.01 (**), p < 0.001 (***), and p < 0.0001 (****). “N.s” indicates no statistical significance.
RESULTS
Adaptive laboratory evolution of novobiocin resistance in S. aureus: genetic mutations and characterization of evolved strains
Through ALE of the ancestral strain (S. aureus ATCC25923), we developed highly resistant strains to NV (Fig 1A and 1B). Whole-genome sequencing revealed mutations in three genes of the resistant strain: gyrB (12/12), potB (5/12), and fpgS (3/12) (S1 Table). The gyrB gene, which encodes the B subunit of DNA gyrase, exhibited non-synonymous mutations across all evolved strains, and therefore conferred high resistance to NV in S. aureus. We selected the evolved resistant strain N10, containing a single gyrB mutation, for detailed biological characterization. The gyrB mutation resulted in slower growth at 37°C than that of the ancestral strain (Fig 1D).
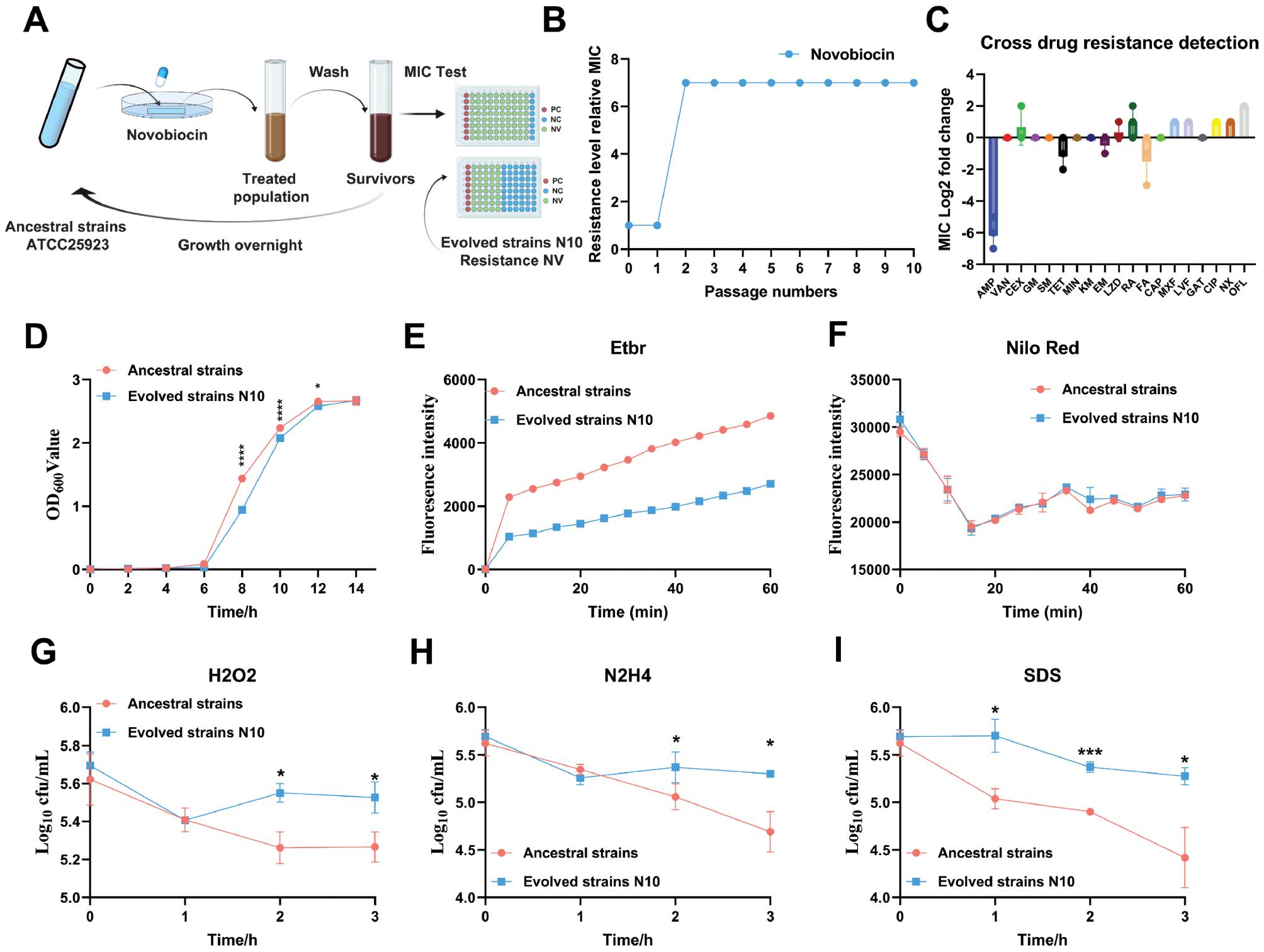
Adaptive laboratory evolution (ALE) of novobiocin in S. aureus and preliminary biological studies of evolved strains.
A. Experimental design of adaptive laboratory evolution. The initial S. aureus strain ATCC 25923, preserved in the laboratory, was cultured overnight in 1 ml TSB at a temperature of 37°C while shaking at a speed of 220 rpm. The culture was then diluted at a 1:50 ratio on the following day and incubated until it reached the stationary phase. A quantity of 200 μl from this culture was coated uniformly onto each TSA plate. In addition, a strip of sterile filter paper measuring 4 cm×1 cm, saturated with an antibiotic solution consisting of a concentration 1 time the Minimum Inhibitory Concentration (MIC). Following a 12 h cultivation period, colonies from the peripheral region of the zone of inhibition were gathered using a disposable loop. The chosen bacteria were wash with 1×PBS and regrow with indicated novobiocin TSB overnight to the stationary phase. The change in novobiocin resistance was measured according to the MIC at each regrowth. This cycle was repeated for ten passages. B. Development of tolerance and resistance in the ATCC 25923 strain under cyclic novobiocin treatment. C. Cross-drug resistance to common anti-Staphylococcus aureus drugs in evolved strain N10. D. Growth curves of evolved strain N10 and ancestral strains. Significance levels are indicated as *p < 0.05; ***p < 0.001; and ****p < 0.0001. E. Mid-log-phase cultures of evolved N10 and ancestral strains were incubated in TSB medium containing 2 μg/mL ethidium bromide for the indicated time. F. Mid-log-phase cultures of evolved N10 and ancestral strains were incubated in TSB medium with 2 μM Nile red stain for the indicated time. G. Mid-log-phase cultures of evolved N10 and ancestral strains were incubated in TSB medium supplemented with 1% H2O2 for the indicated time. The strains were then plated onto TSB plates through serial ten-fold dilution, and the bacteria were counted after 1 day of cultivation. H. Mid-log-phase cultures of evolved N10 and ancestral strains were incubated in TSB medium supplemented with 1% N2H4 for the indicated time. The strains were then plated onto TSB plates through serial ten-fold dilution, and the bacteria were counted after 1 day of cultivation. I. Mid-log-phase cultures of evolved N10 and ancestral strains were incubated in TSB medium supplemented with 0.1% SDS for the indicated time. The strains were then plated onto TSB plates through serial ten-fold dilution, and the bacteria were counted after 1 day of cultivation.
When exposed to 13 common antimicrobial agents for S. aureus, strain N10 demonstrated higher sensitivity to ampicillin, tetracycline, and fusidic acid, but lower sensitivity to cephalexin, linezolid, and rifampin, than the ancestral strain. In contrast, the sensitivity of evolved strain N10 to quinolones, including moxifloxacin, levofloxacin, gatifloxacin, CIP, norfloxacin, and ofloxacin, compared with the ancestral strain, was generally reduced by 2–4 times (Fig 1C). To examine whether altered cell permeability might contribute to altered drug sensitivity in the gyrB mutants, we measured the whole-pore accumulation of Nile red and ethidium bromide, as representative hydrophilic and hydrophobic compounds, through fluorescence spectroscopy. Ethidium bromide accumulated at a slower rate in the evolved strain N10 than the ancestral strain, thus indicating attenuated cell wall permeability (Fig 1E). In contrast, the Nile red accumulation was similar between strains (Fig 1F). Additionally, the survival of the evolved strain N10 and the ancestral strain under oxidative stress was determined. In agreement with the cell permeability results, the evolved strain exhibited enhanced survival under H2O2, N2H4, and SDS stress (Fig 1G–1I). Furthermore, coagulase testing indicated that the evolved strain N10 had diminished clotting ability, requiring approximately 85 ± 0 min for plasma clot formation, compared with just only 65 ± 0 min for the ancestral strain (n = 3, p < 0.05), whereas no statistical significance difference in the morphology between strains was observed on blood agar culture (S2 Table). These data suggested that gyrB mutation alters cell wall permeability and enhances survival under surface and oxidative stress in S. aureus.
Overview of quantitative metabolomics analysis
Metabolic profiling of both the ancestral and evolved N10 strains was conducted with liquid chromatography-mass spectrometry. A principal component analysis model was established to evaluate global metabolite changes in these strains under various conditions both with or without NV. The principal component analysis plots (Fig 2) showed close clustering of samples from each group, indicating consistent metabolic profiles within each group. Notably, the ancestral and evolved N10 strains formed two distinct clusters, thus highlighting substantial metabolic differences between the strains.
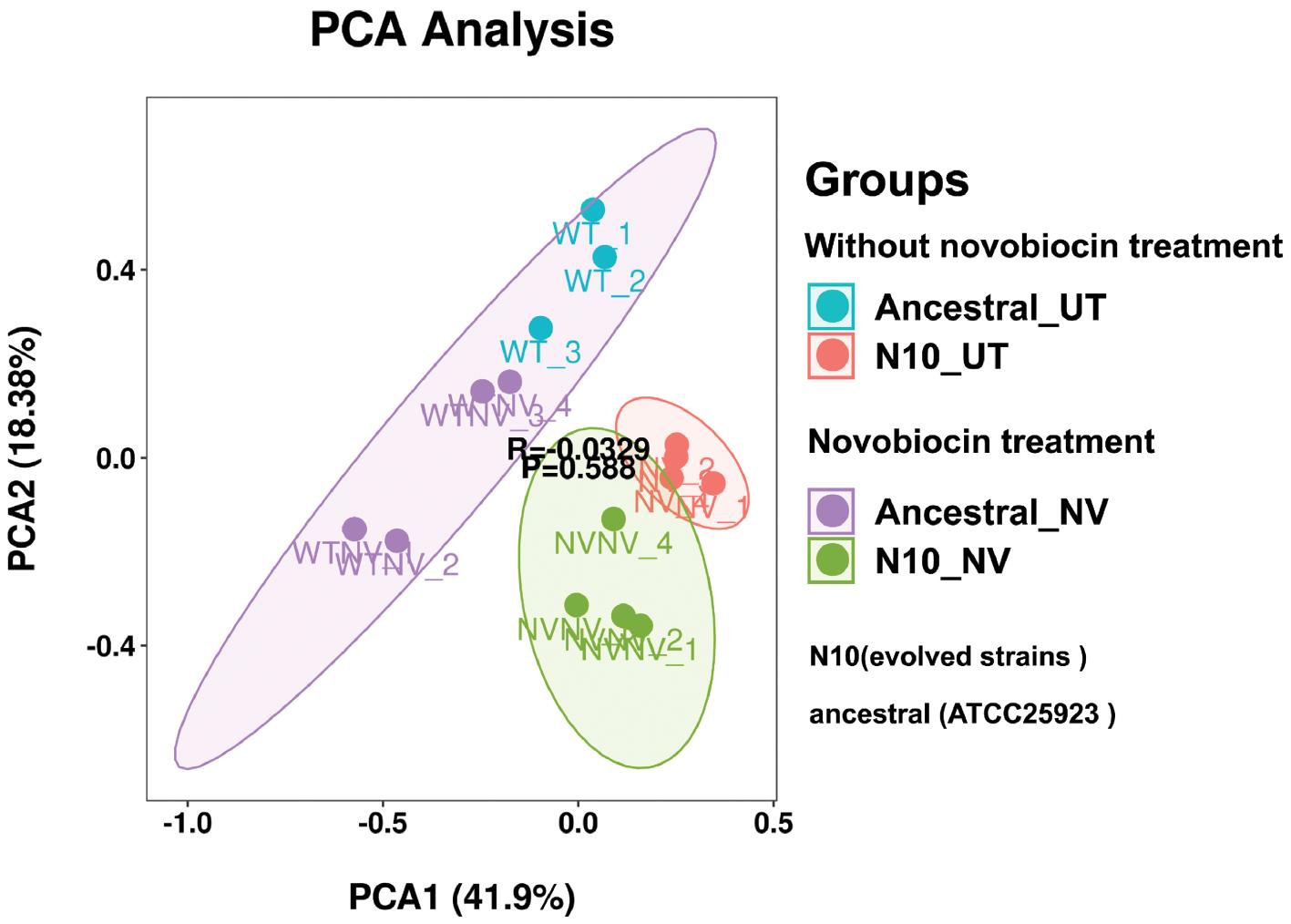
Principal component analysis of metabolites of ancestral and evolved N10 strains before and after novobiocin treatment. N10 represents the gyrB single-gene mutant strain iteratively evolved with novobiocin, ancestral represents the wild strain, UT represents the strain without novobiocin treatment, and NV represents the strain with novobiocin treatment.
The metabolic responses between the ancestral and evolved N10 strains significantly differed, even in the absence of NV treatment, thus indicating that the gyrB gene mutation not only drove NV resistance but also fundamentally altered the metabolic state of S. aureus. Whereas the gyrB mutation was the principal factor conferring resistance, the associated metabolic reprogramming might have contributed to enhanced adaptive capabilities, stress tolerance, and overall fitness of the resistant strain.
Mutation-induced metabolic alterations in evolved S. aureus strain N10
We compared the metabolic profiles of the evolved strain N10 and the ancestral strain to identify changes in metabolite expression induced by mutations. In the untreated N10 strain, compared with the ancestral strain, the levels of 66 metabolites, 56 upregulated and 10 downregulated, showed significant changes (Fig 3A and S3 Table). KEGG and perturbed metabolite network analysis revealed statistical significance upregulation of pathways involved in energy synthesis and amino acid production in the resistant strain, thereby reflecting complex metabolic adaptations to genetic mutations and pharmacological pressure (Fig 3C and 3D).
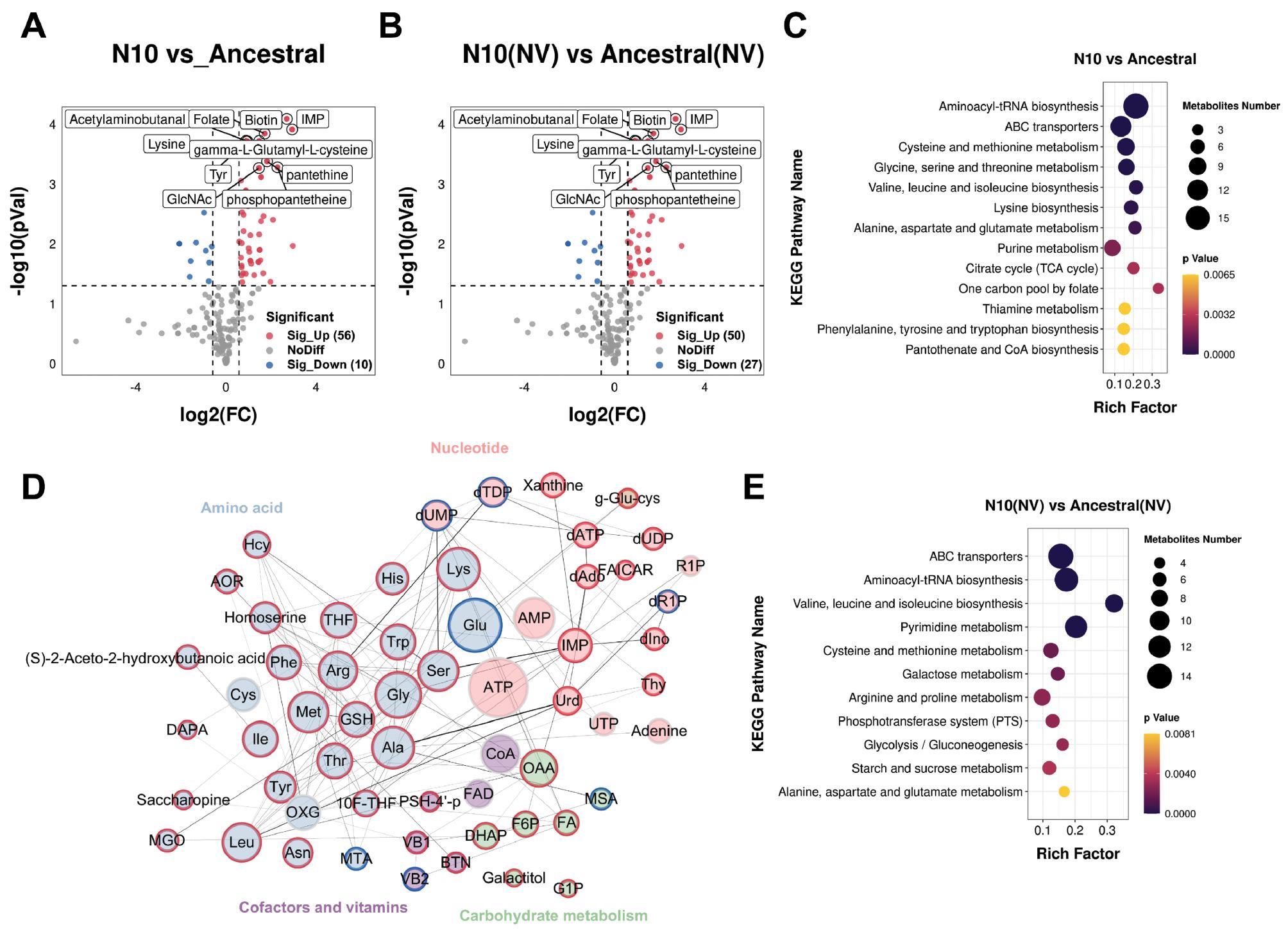
Overlapping and specific metabolomic responses between ancestral and evolved N10 strains, with or without novobiocin treatment. N10 represents the gyrB single-gene mutant strain iteratively evolved with novobiocin, ancestral represents the wild strain, UT represents the strain without novobiocin treatment, and NV represents the strain with novobiocin treatment.
A and B. Volcano plots for evolved strain N10 compared with the ancestral strains in the absence of NV (A) and after NV treatment (B). C. KEGG enrichment analysis of differentially expressed metabolites (DEMs) in evolved strain N10 compared with ancestral strains in the absence of novobiocin. D. Network analysis of DEMs in evolved strain N10 compared with the ancestral strain without NV treatment. Lines represent metabolite interactions, with thicker lines indicating higher confidence levels. Larger circles indicate higher degrees of interaction. The colors inside the circles represent different metabolic pathways. The outer circle border is blue for down-regulated metabolites and red for up-regulated metabolites. E. KEGG enrichment analysis of differentially expressed metabolites (DEMs) in evolved strain N10 compared with ancestral strains after novobiocin treatment.
After NV treatment, 46 metabolites, 32 upregulated and 14 downregulated, exhibited statistical significance changes in the evolved strain N10 compared with the ancestral strain (Fig 3B and S3 Table). KEGG analyses revealed statistical significance upregulation in pathways including amino acid biosynthesis, ABC transporters, starch and sucrose metabolism, and glycolysis/gluconeogenesis pathways. Specifically, upregulated metabolites associated with amino acid biosynthesis included serine, threonine, leucine, isoleucine, methionine, arginine, lysine, and histidine. Additionally, pyrimidine-associated metabolites were downregulated, whereas purine-associated metabolites were upregulated in the N10 strain compared with the treated ancestral strain after NV treatment. These changes in metabolite levels suggested that the evolved N10 strain exhibited altered metabolic responses with NV treatment, thus potentially reflecting adaptations supporting its resistance phenotype (Fig 3E).
Metabolic responses to novobiocin treatment in ancestral and evolved S. aureus strains
After NV treatment, compared with no treatment, 37 metabolites exhibited statistical significance changes in the ancestral strain. Specifically, 29 metabolites were upregulated, whereas eight metabolites were downregulated (Fig 4A and S3 Table). The upregulated metabolites were associated predominantly with purine and pyrimidine metabolism, including compounds such as CAIR, FAICAR, deoxyinosine, CDP, CTP, and UDP. Similarly, metabolites associated with amino acid biosynthesis, such as glutamate, glutamine, acetylornithine, citrulline, GABA, and histidine, also showed increases (Fig 4C and 4D). Additionally, metabolites involved in pantothenate and CoA biosynthesis, such as coenzyme A, cis-2-methylaconitate, and dephospho-coenzyme A were elevated. The downregulated metabolites were involved primarily in glycolysis, and included (S)-3-methyl-2-oxopentanoic acid, dephospho-coenzyme A, 2-phospho-D-glycerate, dimethylmalate, guanosine, phosphoenolpyruvic acid, and 3-phosphoglyceric acid (Fig 4C and 4D).
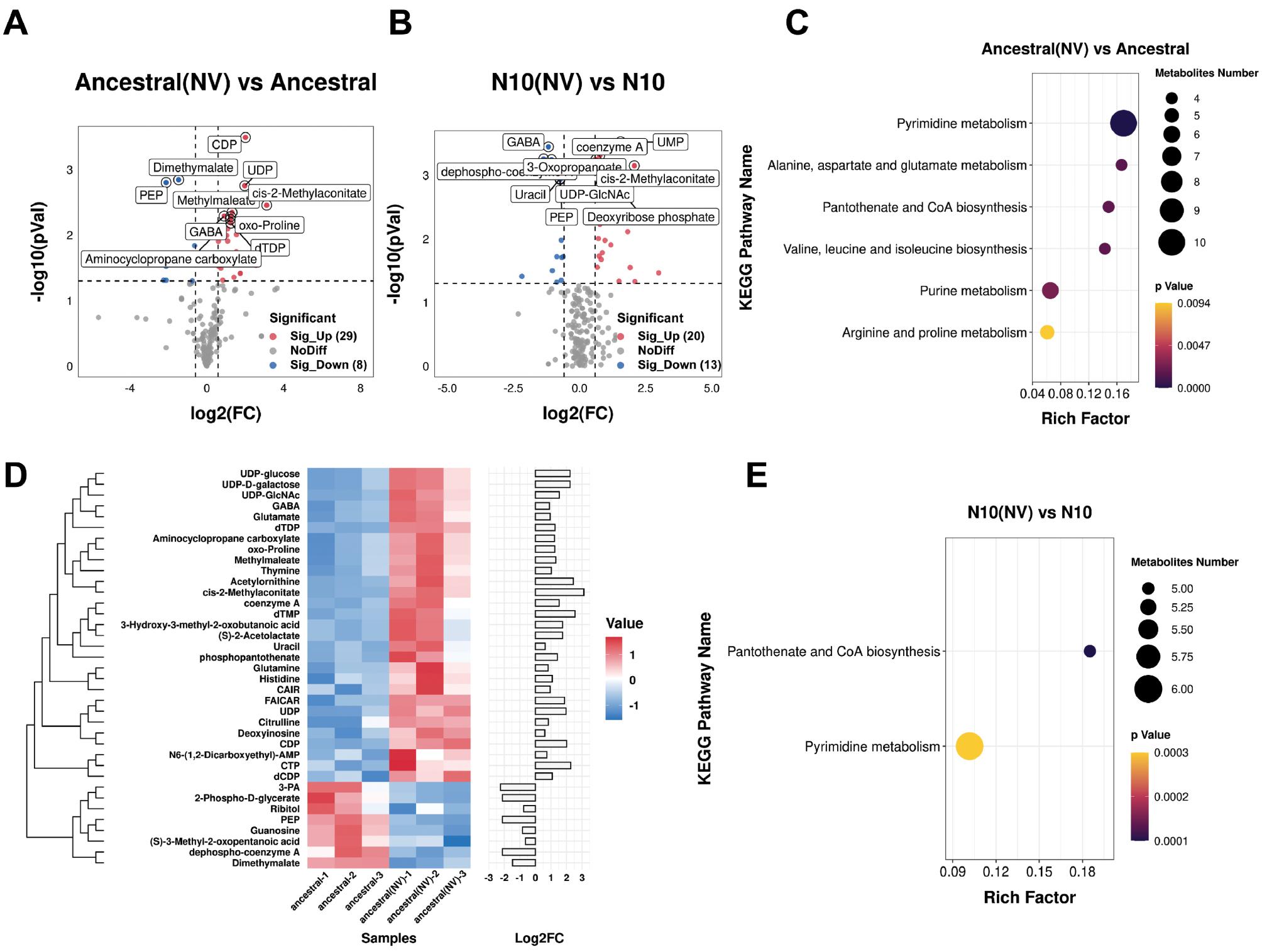
Overlapping and specific metabolomic responses to novobiocin between ancestral and evolved N10 strains. N10 represents the gyrB single-gene mutant strain iteratively evolved with novobiocin, ancestral represents the wild strain, UT represents the strain without novobiocin treatment, and NV represents the strain with novobiocin treatment.
A and B. Volcano plots for evolved strain N10 and ancestral strain compared with each strain without NV treatment. C. KEGG enrichment analysis of differentially expressed metabolites (DEMs) in the ancestral strain after NV treatment versus without NV treatment. D. Heat map of DEMs in the ancestral strain after NV treatment versus without NV treatment. E. KEGG enrichment analysis of differentially expressed metabolites (DEMs) in evolved strain N10 after NV treatment versus without NV treatment.
In the evolved N10 strain treated with NV, compared with no treatment, statistical significance changes were observed in 33 metabolites, 20 upregulated and 13 downregulated (Fig 4B and S3 Table). The upregulated metabolites included primarily pyrimidine metabolism products such as CDP, CTP, dTMP, UDP, and UMP, as well as purine metabolism intermediates, including CAIR, dAMP, deoxyadenosine, FGAM, and GTP. Both the evolved N10 strain and the ancestral strain exhibited 17 DEMs after NV treatment, affecting primarily pyrimidine metabolism, pantothenate and CoA biosynthesis, and glycolysis (Fig 4E). These findings suggested that these pathways might respond to NV stress, despite not being directly associated with NV’s mode of action (MoA). Furthermore, after NV treatment, amino acid metabolic intermediates (S-3-methyl-2-oxopentanoic acid) and purine synthesis intermediates (CAIR) were upregulated in the evolved N10 strain but downregulated in the ancestral strain. These findings highlighted distinct adaptive and response strategies between strains at the genetic or regulatory level (Fig 4C and 4E).
DISCUSSION
Drug phenotypes represent complex, higher-order functions that, although clearly defined through experimentation, result from a cascade of intricate mechanistic events, each initiated by a specific molecular interaction. Traditionally, drug development and MoA studies have focused on the initiation of these interactions, primarily because of their chemical tractability and the assumption that the subsequent stages of MoA lack clear specificity. However, the emergence of systems-level technologies has increasingly challenged this perspective. The identification of precise system-level indicators, or signatures, of drug efficacy has transformed understanding of drug activity, by offering profound insights into both primary and secondary effects. Consequently, secondary or downstream MoA-related responses have become powerful tools for classifying drug actions, even in the absence of direct knowledge of their targeted molecules. This shift highlights the importance of comprehensive systems-level approaches in contemporary drug research. Metabolomic studies comparing NV-resistant and NV-sensitive S. aureus can uncover resistance mechanisms and guide treatment strategies [20].
ALE and whole-genome sequencing demonstrated that S. aureus developed NV resistance through gyrB mutation, thus decreasing growth rates and coagulase activity. Mutations in potB and fpgS, involved in polyamine transport and the folate pathway, respectively, might also contribute to this phenomenon [21,22]. Future studies should use a genetic editing approach to verify the contributions of these mutations to NV resistance. Furthermore, we observed that resistant strain exhibited enhanced sensitivity to antibiotics such as ampicillin, fusidic acid, and tetracycline, which affect protein synthesis and cell wall synthesis. When DNA gyrase function was impaired by gyrB mutations, the bacteria’s ability to manage these biological processes was compromised, and the susceptibility to these antibiotics increased. In contrast, resistant strain exhibited increased tolerance to cephalexin, linezolid, and rifampicin, thus suggesting that the effects of gyrB mutations on other non-DNA gyrase targeting drugs varied. This variability might have stemmed from gyrB mutations altering the targets of these antibiotics or influencing downstream antibiotic processes. Furthermore, chromosomal mutations leading to antibiotic resistance are well established to frequently alter essential targets such as ribosomes, DNA gyrase, RNA polymerase, or cell wall synthesis [23–25]. Membrane permeability assays and surface and oxidative stress results showed that the gyrB mutation decreased membrane permeability and enhanced survival of S. aureus under in vitro stress conditions. These target changes often impair critical functions or impose metabolic burdens, thus resulting in fitness costs and detrimental effects [26–28].
Our metabolomic analysis of ancestral and evolved N10 strains (gyrB mutant), with or without NV treatment, revealed significant shifts in bacterial metabolism, attributable to both the pharmacological effects of the drug and intrinsic metabolic reprogramming due to the gyrB mutation. In the ancestral strain, upregulation of purine and pyrimidine metabolic pathways was likely to serve as a compensatory mechanism under NV stress. Impaired DNA replication and transcription led to decreased availability of raw materials for energy synthesis, thereby increasing cellular energy demand. In addition, amino acid biosynthesis pathways, particularly those for isoleucine, alanine, and arginine, were significantly upregulated. Branched chain amino acids in S. aureus are crucial factors determining its virulence and drug resistance, with changes in virulence and drug resistance being mainly associated with the utilization of isoleucine [29,30]. Similarly, the synthesis and utilization of alanine were also associated with S. aureus virulence and survival under antibiotic stress. Disruption of alanine utilization leads to bacterial malnutrition and impaired survival [31,32]. S. aureus is the only known bacterium capable of synthesizing arginine from proline through the arginine-proline interconversion pathway, a process that counteracts host carbon metabolism and is critical for bacterial infection and antibiotic tolerance [33,34]. In addition, compensatory upregulation of virulence-associated amino acids can trigger the stringent response, a regulatory mechanism that helps bacteria cope with stress. Although antibiotic sensitivity can widely vary, the general resistance to quinolones observed in S. aureus might have resulted from the stringent response triggered by global metabolic reprogramming after gyrB mutation.
Furthermore, the upregulation of pathways enhancing the synthesis of virulence amino acids and mitigating drug-induced damage similar to the recent reports. A previous report on Salmonella has indicated that damage to DNA gyrase leads to decreased expression of the virulence factor SPI-2, thereby impairing bacterial survival within macrophages [35]. Although the above hypothesis requires further validation, differential mechanisms were observed between strains with varying sensitivity to the same drug, as also observed in vancomycin-intermediate and vancomycin-resistant S. aureus (VISA and VRSA) [36]. Importantly, metabolites associated with the TCA cycle, CoA metabolism, amino acid metabolism, and nucleotide metabolism were found to be significantly altered. These findings are consistent with those from previous reports of antibiotic-induced metabolic changes in other bacteria [11,37]. In a related study [38], the metabolic responses of MRSA and methicillin-sensitive S. aureus to ampicillin, kanamycin, and norfloxacin have been investigated. Whereas ampicillin and kanamycin caused pronounced disruptions in metabolites associated with pyrimidine and purine metabolism, these pathways showed only moderate to low levels of disruption after norfloxacin treatment. These findings indicate that, although norfloxacin is a quinolone that targets bacterial DNA synthesis, NV might exert a stronger effect on bacterial metabolism. This observation aligns with findings that a topoisomerase IV mutation in Escherichia coli requires a DNA gyrase mutation as a prerequisite for effective metabolic disturbance [39].
After NV treatment, the DEMs shared between the evolved N10 strain and the ancestral strain influenced predominantly pyrimidine metabolism, pantothenic acid and coenzyme A biosynthesis, and glycolysis. This correlation was expected, because previous studies have demonstrated that gyrA expression is induced by overexpression of ArcA, a global respiratory metabolic transcription factor in E. coli, thus highlighting the regulatory effects on DNA replication and metabolic state changes, particularly in glycolysis [40]. Moreover, the aggregation of S. aureus is essential for its virulence, and the glycolytic pathway plays a critical role [41]. As previously described, the fitness cost imposed by gyrB mutations might lead to compromised virulence. Thus, we propose that the upregulation of glycolysis caused by gyrB mutations is necessary to compensate for the loss of virulence. Similarly, the pantothenate pathway has also been demonstrated to be associated with virulence and drug resistance [42,43]. Therefore, upregulation of the pantothenate pathway might serve as a potential mechanism to compensate for virulence and confer cross-resistance.
Bacterial metabolism is intricately associated with antibiotic sensitivity [44,45]. Notably, despite the presence of the DNA gyrase inhibitor NV, the evolved N10 strain exhibited significant upregulation of key metabolic pathways, including amino acid biosynthesis, ABC transporters, starch and sucrose metabolism, the glycolysis/gluconeogenesis pathway, and purine and coenzyme pathways. These changes are believed to be compensatory responses to the gyrB mutations in S. aureus. DNA gyrase inhibitors such as quinolone or norfloxacin have been shown to induce cellular gyrase poisoning, which leads to the production of highly damaging hydroxyl radicals via the Fenton reaction [40,46]. Subsequently, a maladaptive response exacerbates the primary effects of gyrase inhibition, by causing oxidative damage to DNA, proteins, and lipids [47–50].
Our findings illustrate that even drugs such as NV, which target non-metabolic processes, provoke significant metabolic alterations in bacterial systems—such as in protein synthesis and DNA replication—following critical genetic mutations that confer resistance. These adaptations often represent compensatory adjustments by bacteria to mitigate drug pressure, either by reallocating energy or facilitating metabolic bypass. Despite these fitness costs, evidenced by impaired growth and coagulation, resistant strains also exhibited increased resistance to quinolones, enhanced membrane impermeability, and improved survival under oxidative stress—phenomena similarly observed in fusidic acid-resistant S. aureus [51]. Importantly, our results emphasize the potential metabolic role of the gyrB subunit in mediating responses to gyrase inhibitors, and reveal a previously underexplored link between gyrB mutations and bacterial metabolism.
In conclusion, through ALE, we developed S. aureus exhibiting resistance to NV. Metabolic analyses of strains with single-point mutations in the gyrB gene revealed that these resistant strains underwent compensatory metabolic adjustments leading to significant alterations in crucial cellular processes, such as protein synthesis and DNA replication. These findings underscore gyrB’s critical role in bacterial resistance and provide previously uncharacterized insights into the metabolic underpinnings of antibiotic resistance.
Our study further highlights the importance of targeting bacterial metabolic pathways as a complementary strategy to combat resistance. For example, therapeutic approaches combining traditional antibiotics with inhibitors of key metabolic pathways identified in this study (e.g., purine, pyrimidine, or amino acid biosynthesis) might offer novel treatment avenues. Further research, including functional validation of these pathways and the development of targeted metabolic inhibitors, is critical to enhancing antibiotic efficacy. Additionally, investigating the evolution and dissemination of drug resistance across bacterial species through multi-omics approaches might enable generalization of these findings and contribute to predictive models integrating genetic, metabolic, and phenotypic data. Such models would accelerate the discovery of effective antimicrobial agents against emerging resistance mechanisms.


